Before getting into this week’s post, I’d like to remind my readers that the discoveries of plant science I talk about here are going on in plants everywhere. The details may vary and the specific output responses might be different, but the research plant scientists do translates to most plants. The processes that go on inside the plant are often similar. It’s something to think about as you walk outside and see the green world.
Doing science
If you’re a scientist, most experiments don’t yield amazing and pathbreaking results. They tend to be small pieces to a larger puzzle. Or they are trouble shooting, quality control, or checking to see if a previous hypothesis was wrong by seeing if its predictions hold up. Scientific stories are the hypotheses that stand up to scrutiny or the refutation of hypotheses that don’t. That scrutiny starts with the scientist doing the work. Next, it gets written up and goes through the peer review system. When published, the research is there for other scientists to see/review and use to inform their own work. Publications can be fodder for scientists to talk to one another in the continuing endeavor to describe nature as best we can. Or work on ways to apply what we have learned to solve a problem in the world with private sector scientists. Progress happens, but the story is never complete.
A lot of the time, this is a slog. Scientists try to design experiments that will yield informative findings, but a priori, we don’t know what will work and what won’t. It’s not that there aren’t good moments, but they’re often small. This post is about what I consider my best moment in contributing to science, where I felt proud about what I’d uncovered.
Scientists try to design experiments that will yield informative findings, but a priori, we don’t know what will work and what won’t.
What was your best moment in science? Or if you’re reading this and aren’t a scientist, what is a moment you’ve had where you felt onto something or understood something for the first time. Where possibilities opened up. Think about it and feel free to share it in the comments (even if you skip the rest of this post; I won’t take offense, I swear).
The backdrop to my best moment in science, getting sob3-6.
During my Ph.D. I was studying some odd genes that are unique to plants. They have one part that can bind to DNA, a so-called AT-hook. This means they can bind Adenine-Thiamine-rich sequences of DNA. This suggests they are transcription factors, proteins that can affect expression of other genes. And they had another part, a “domain of unknown function” in the parlance of protein databases (it was dubbed a Prokaryote-Plant Conserved– PPC– domain in one publication). It’s a sequence that comes up many times in genome databases, but had no known function during my Ph.D. In Arabidopsis (the lab-rat of the plant world), there are about 30 members of this protein family encoded in the genome. This isn’t uncommon in plants or in biology to have many related genes in a single genome. In fact, many plants have whole genome duplication events in their evolutionary pasts. This can often mean that the genes overlap in function. They may even be redundant, making them somewhat recalcitrant to genetic studies. A common way to study things in genetics is to break genes and see what happens to the organism. If two genes go the same job, you can cross broken (mutant) versions of them together to create a double mutant. These “higher-order” mutants take a fair bit of work to create. And even more difficult if two genes are located close together in the genome (more on a version of this problem later).

The gene I worked on in my Ph.D. is SOB3. SOB stands for “suppressor of phytochrome B” (& yes, there are puns you can make with the name SOB). phytochrome B (phyB for short) is one of the plants informational light sensors. Phytochromes recognize the red/far-red spectrum of light, and if phyB function is lost, plants become spindly and elongated. They are always looking for an optimal light environment but can’t find one because they are blind. One of the responses that is easy to look at is the seedling response. The seedling stem (the hypocotyl in plant anatomy lingo) stops elongating in response to light. In a phyB loss-of-function plant, hypocotyls are not suppressed and elongate more. It is akin to what a seed that germinates in the dark does (think about bean sprouts you put on your sandwich).
Look at the Quiet Branches cover photo and that one seedling that is elongating more and is white/paler with closed leaves. Most plants have more than one phytochrome. Arabidopsis has 5. As well as other light sensors for other parts of the light spectrum. They are redundant in some ways. phyB, however, seems to have the largest affect on plant growth and development. SOB3 was found in a mutant screen where the goal was to induce over-expression of genes that could suppress the physical look of a partial loss-of-function version of phyB (a plant that still made phyB, but one that didn’t work as well as the normal version). In other words, to find plants that still had little phyB function, but looked like they were normal again. The screen was designed to dial up expression of random genes in the genome with the goal of letting the plant tell the scientists what is important. I won’t go into the details but the approach involved making transgenic plants. Inserting a foreign piece of DNA and landing that inserted piece of DNA near a gene that would dial up it’s expression. It’s pretty clever and is called activation tagging. And if that gene had something to do with red-light signaling, it would revert a phyB mutant to a normalish looking seedling.
Over-expressing SOB3 suppressed the phyB mutant phenotype. In fact, it made the suppression even more severe than normal resulting in an even shorter hypocotyl. My best moment was finding that one on the far right:
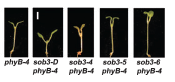
Another consequence of over-expressing SOB3 was a rather dramatic adult phenotype (how something looks/appears). The plants were gigantic and long lived. They produced gigantic leaves, thicker stems and bigger seeds, my Ph.D. advisor nicknamed it “Shaq”. Two talented undergrads actually cloned the gene (identified what it was as described above). Another gene similar to SOB3 was also identified in a similar approach. In fact it’s the next closest gene in sequence similarity to SOB3, dubbed ESCAROLA (ESC) due to the adult look of the plant that made it look like escarole lettuce. So I took that one on too in my project of figuring out just what these genes actually did. As I noted in my last post, it’s essential to be careful when over-expressing genes and interpreting the results. Off-target effects can occur or the whole system of the plant may be perturbed and obscures what a gene does. The best thing to get is a loss-of-function gene that has the opposite phenotype of the over-expressor. That is at least promising that you have zeroed in on the process(es) which a gene (or family of genes) is involved. A big part of my project was to identify loss-of-function versions (alleles) of SOB3 and ESC…and hopefully find a phenotype. I did not want to have to knock out all 30 of the family to see an effect! With modern techniques, that may now be at the edge of possibility. But it is still a lot of work to identify and cross together all the different loss-of-function genes into a single plant. This is why breakthroughs in plant biology, desired agronomic traits, take awhile to make it into farmers hands. The traits have to be bred into varieties known as “elite varieties” of plants that farmers grow with other desired traits. Genetics takes time and generations (and so many scientists don’t like it. Faster is better).
Genetics takes time and generations
I remember hearing years ago about the identification of a naturally decaffeinated coffee tree. It’s a variety of coffee that does not make caffeine! That’s amazing, but scientists have to breed that variety into commercial coffee trees before it can farmers can grow it and get it out to the world. When that happens, decaf will no longer be produced by filtering caffeine out through chemical processes. Those processes work, but to my taste budes, make de-caf coffee taste terrible. The crossing can take a long time. If you ever wondered why innovation takes time, this is one reason why; bringing the innovation into a form for market release.
Losing SOB3 and ESC function
As it is in agriculture, so it is in the lab. In my case, it was to identify loss-of-function versions of SOB3 and ESC– to break these genes. ESC was easy. Someone had generated a series of point mutations through the ingenious TILLING project. It was a targeted way of finding changes in DNA in your gene of interest. One of those mutations caused an early stop, so neither of the conserved domains would get translated into protein. SOB3 was a bit harder. The T-DNA insertion database had a few insertions that were near the gene that can sometimes disrupt function. One was just 3 base pairs from where the protein coding sequence of the SOB3 gene would start. A good sign, but it also seemed to overproduce the RNA transcript…not eliminate it. However, it did not have the SOB3 over-expression phenotype, so it was put into the “may be loss-of-function pile”.
My Ph.D. advisor had a very clever suggestion. It was to take the original SOB3 over-expressing line and treat it with a chemical that induces changes in DNA (aka mutations). And then look for plants that had lost the over-expression phenotype. Sequencing the SOB3 gene in these mutated lines might reveal changes in SOB3 rendering it non-functional. And I found one. It was similar to the esc mutation, but in SOB3. It was an early stop so no functional protein would be made (though it was still over-expressing the SOB3 transcript..it was just not functional anymore). I crossed both my “maybe” and my early stop sob3 mutants to esc. In genetics, mutant versions of genes are written in lower case italics. I got sob3 esc double mutants (that is individual pants carrying the loss-of-function versions of SOB3 and ESC) and compared them to normal plants. They had a subtle, but detectable, hypocotyl (seedling) phenotype opposite that of the over-expressor. Subtle phenotypes may indeed be real. But they can be misleading, may disappear on further measurement or under different conditions, and more. Luckily, I had uncovered two other mutations from this intragenic suppressor screen.
Curse you, Genetics!
Both of these other suppressors were near or in the AT-hook DNA binding domain of the protein. They change the amino-acid make up of the protein, but don’t prevent it from being made by the plant. These are called missense mutations. One was nearby the core 3 DNA binding amino acids and it had a long hypocotyl as a single mutant. The other, sob3-6, had a mutation in the central amino acid of the DNA-binding domain. Bullseye. However, to get at it, we were quite lucky. I had a great undergrad working with me on this screening. From the screen, there were 4 plants that looked like they had suppressed the over-expression phenotype, 3 of the 4 had what is known as a pin-formed phenotype:

This phenotype is what happens when auxin transport is defective (PIN1 is a gene that exports auxin from cell). These mutants are not viable. You can force them to produce some flowers by treating them with auxin (and I did that!), but the flowers were not normal flowers and they yielded no seeds. There was the one plant that looked pretty normal though and did produce seeds.
There was the one plant that looked pretty normal though and did produce seeds
We sequenced all of these 4 plants and found the mutation we called sob3-6. The sequence results were quite obvious for the 3 pin-formed mutants. Had this mutation in DNA binding domain of SOB3 caused it?
When the undergrad sequenced the 4th plant, the one that looked normal, he though initially it didn’t have the sob3-6 mutation. I went back and took a closer look and noted a ‘double peak’ where the sob3-6 mutation was; it looked like there were two different base-pairs where only one should have been.
What’s neat and a pain for scientists in complex organisms like humans and plants is that we have two copies of our genes; one from Mom and one from Dad. Arabidopsis is a self-fertilizing plant, and further experiments revealed that that 4th plant, the normal looking one, had one mutant (i.e. sob3-6) copy and one normal copy of SOB3– the genetics word for this is heterozygous (as opposed to homozygous).
In the next generation, it segregated. If you remember learning genetics in high school about Mendel and his peas, the prediction is you’ll get a specific ratio of plants in the next generation. Namely, 1 homozygous for the normal SOB3 (though remember it’s still the over-expressor still), 2 heterozygotes, and 1 homozygous for the sob3-6 mutation. Whenever sob3-6 was homozygous, it was also pin-formed. It also had an extreme in length hypocotyl. Did sob3-6 cause both hypocotyl and pin-formed phenotypes?

Thanks to the Arabidopsis genome project, The chromosome containing SOB3 had it’s complete sequence known and annotated. Nearby SOB3 was the PIN1 gene, an auxin transporter known to cause a pin-formed phenotype when its function is lost. What was likely happening was that my sob3-6 version was being inherited in the next generation along with the annoying pin-formed mutation. The mutating chemical I used is random; it causes several thousand mutations in a genome all at once. So I had a new mission: Break the linkage between these two mutations. This involved planting a lot of the segregating sob3-6 heterozygous seeds and growing enough to catch a chromosomal rearrangement. Through normal biology, there’s a part of the reproductive cell cycle (i.e. meiosis) where the two copies of chromosomes from each parent can exchange DNA with their sister chromosome. Barbara McClintock showed this with her experiments in Maize in the 1930’s.
I had to find the plant where recombination happened. That is, the plant where a non-pin-formed chromosome recombined with a pin-formed one. Thus leaving behind a functional PIN1 gene in combination with my sob3-6 mutation. And I found it! After planting lots of plants and a lot of genotyping.
sob3-6 does have a long hypocotyl. Even when it is heterozygous (present as only one copy in the genome), it had that phenotype. Again, if you think back to Mendel and his peas, he came up with the idea that some traits were “dominant” over others; they showed up in how an organism looked when present in one copy. sob3-6 appeared to be dominant. But in a specific way. It caused an extreme phenotype, but the mutation had to be present to see it. It appeared to be something called a ‘dominant negative’ mutation. I think of it as a mutation that gums up the works of normal cellular machinery. The full protein is made in sob3-6, but it presumably can no longer bind DNA. So if it’s needed as part of gene regulatory machinery involving other proteins and it’s lost an anchor to DNA, several proteins are now just floating around the cell bound to the sob3-6 protein, not doing their normal job. And because it was being over-expressed, it could wreak even more havoc for that reason too.
Triumph!
Identifying sob3-6 is my proudest moment in science. Up to the point of my paper on SOB3 (previously named AHL29), the gene family was relatively uncharacterized. I could count the number of publications about it on one hand. So even if it’s small, I feel I moved the field of study forward.
As far as I found, the only thing sob3-6 had an effect on was the seedling in the hypocotyl. But making similar mutations in some of the 29 other members of the family might show different effects. All depending on where and when a SOB3 family member is expressed during development.
My Ph.D. advisor has carried on working on SOB3 and the rest of that gene family. I am not sure if they are the next great innovation in plant science. It’s probably too soon to know. But new evidence from my Ph.D. advisor’s lab suggests that these proteins interact with each other through that domain of unknown function in groups of 3 as well as with other transcription factors (Zhao et. al., 2013). It makes sense in light of the sob3-6 phenotype. It’s dragging away binding partners from DNA where they’d normally act.
Perhaps targeting them for gain- or loss-of-function can alter gene expression in important ways in agriculture. If you can target the expression of sob3-6, it may be able to confer a desirable trait to a plant. The only thing I don’t know is if you express sob3-6 at a normal level, if it still has a dominant negative phenotype. It may not matter. In the end, it’s just a really interesting story. And one that required persistence, luck, awareness, and team-work.
sob3-6 was a cool moment. I’d helped confirm my complete loss-of-function sob3 esc data. And I identified a really cool manipulation of a plant through change in a single base pair (OK, fine, with some over-expression too).
Share your story in the comments. Or just email it to me via the contact form below and I may publish some of them in a future post with your permission, of course.
References

One thought on “My greatest moment in plant science.”