
Cells are the units that make up life. Multi-cellular organisms like humans or plants can be millions or trillions of cells. Most cells are also specialized in function. Skin cells look and do very different things from a brain neuron. A leaf cell is much different than a root cell. Specialized (i.e. differentiated) cells started as undifferentiated stem cells.
How does a cell figure out its identity? That is a big topic in developmental biology.
How does a cell figure out its identity? That is a big topic in developmental biology.
Two recent papers in The Plant Cell on Arabidopsis root development provide some answers to this question. The broad answer is that plants (most living things, actually), have genetic programs it runs that define the cells that make up the root.
From a seed
Inside a plant seed is an embryo. For this story, the relevant part is the few cells that will grow into the entire root of the plant. After germination, these few cells that are the new root start dividing and establish a root designed to find water, soil nutrients, interact with fungi/microbes in the soil, and anchor the shoot system (that has its own meristem and story for another day). After a population of dividing cells (also known as transit activated cells or TACs) is established, elongation and differentiation begin to occur. A dynamic balance of cell division and differentiation is then in place with the in-between territory (where a cell isn’t fully differentiated, but unlikely to divide) being the transition zone.
Depending on conditions, cell division activity can increase, differentiation can increase, or both can be in equal balance.
A population of stem cells is maintained throughout root growth as plant growth and development occurs almost entirely post-embryonically– post seed– a contrast with animals where a lot of development happens before birth.
This is the root meristem. A constantly maintained population of stem cells, actively dividing cells. The meristem then ends and cells transition to elongation/differentiation of cells occurs.
How does a root cell know if it’s a stem cell? A dividing cell? A differentiating cell? Two recent papers in The Plant Cell address this question. As cells move from dividing stem cells (that divide to maintain a stem cell and a cell that will become a transit activated cell) through the meristem, and into differentiation, there are specific genetic programs that define each area and let the cells know their identity. As cells move, differing instructions cause cells to adopt their new behavior/function. This can occur across a single cell distance, as the dividing stem cell example in this paragraph shows.
How cells know who they are
Plant cells are constrained by a rigid cell wall and don’t move. Part of how a plant cell knows what to do is based on its position. In the root, a key factor is distance from the stem cell niche (SCN). Another is the internal-external root axis (aka across) consisting of files/rows of cells. Roots mostly have radial symmetry, though some of the vascular tissues, near the center of the root, adopt a two-sided (bilateral) symmetry.
How does a plant cell tell it’s position and what it’s supposed to do at any given time?
Stem Cells vs. Dividing Cells

Rodriguez et al. (1) provide evidence for one mechanism by which plants differentiate stem cells vs. the zone of cell division. The researchers looked at a gene regulatory loop that helps define the stem cell niche and demonstrated how it is altered when the scientists manipulated either one or the other of these components. One of these components is a family of transcription factors (proteins- gene products- that turn other genes up or down), dubbed “GROWTH-REGULATING FACTORS“, or GRFs. The other is a microRNA, specifically microRNA396, or miR396. microRNAs are genes encoded in the DNA of the genome. However, the gene product is a small (21-24 nucleotide) RNA molecule of specific sequence that can prevent the making of proteins whose RNA complements the microRNA’s sequence (creating a double stranded RNA that cells can cut apart). microRNAs negatively regulate genes. In the case of miR396, it’s target is the GRF transcription factor family.
The PLT-mir396-GRF genetic circuit is just one of many operating, but it is one way plants specify identity.
Rodriguez et al. found that that mir396 is specifically expressed in the stem cell niche and keeps the GRFs from being expressed there, maintaining stem cell identity. The GRFs, expressed in the meristem/TAC cells, at least partially defines those cells as actively dividing. GRFs repress a family of transcription factors that also define stem cells, the PLETHORA (PLT) genes. PLT actually turns on miR396 in the stem cell niche. As a cell leaves the stem cell niche, PLT expression is repressed by the GRFs, promoting transit-activated status. The PLT-mir396-GRF genetic circuit is just one of many operating, but it is one way plants specify identity.
Cycling vs. non-cycling cells
Otero et al. (2), the scientists looked at a rather generic type of protein: histones. Histones are proteins that are part of packaging DNA in cells. Histone octamers (groups of 8) have DNA wound around them, a bit like spools of thread. Each octamer has around 150bp of DNA wrapped around it. This means that per chromosome (thousands-millions of base pairs), there are a lot of histone core complexes. There are also linker histones, that bind DNA that spans the distance between histone core octamers. Histone octamers can be loosely or tightly packed together or moved along the DNA, making parts of the genome more or less accessible to things like the above mentioned transcription factors. This matters for what genes will be active or not in any given cell.
Histone octamers can be loosely or tightly packed together or moved along the DNA, making parts of the genome more or less accessible to things like the above mentioned transcription factors. This matters for what genes will be active or not in any given cell.
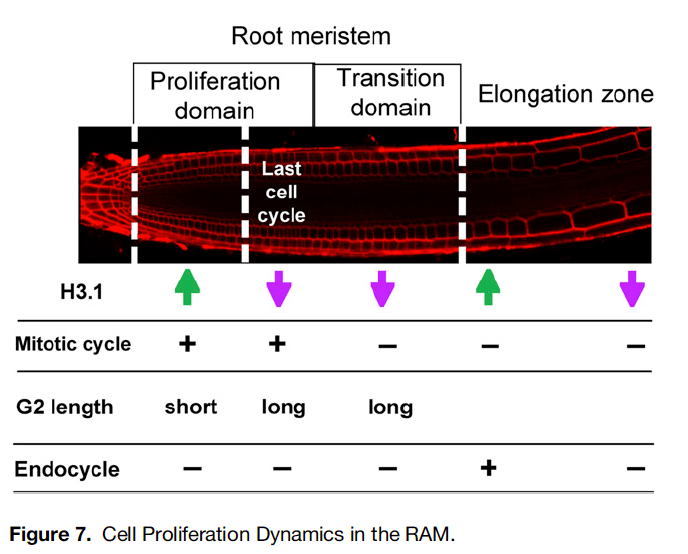
There are variants of histones in the genome, or several different types comprising the octamer. One component of the octamer are histones of the family H3. Specifically, Otero et al. looked at tagged versions of H3.1 and H3.3 and looked at how H3.1 in particular gets replaced from histone octamers at some specific times in root cell development. They noticed that a high ratio of H3.1/H3.3 seems to mark cells for high proliferation ability (starting with the center of the stem cell niche, the Quiescent Center, that contains cells that do not divide, and are therefore lacking in H3.1, but have H3.3). H3.1 seems to get replaced from DNA when cell division is going to cease and the scientists demonstrated this for the root meristem. Not only does a high H3.1/H3.3 mark cells as actively proliferating and going through the cell cycle, when cells approach the transition zone/cessation of cell division, the final cell division has lengthier G2 cell cycle phase, likely because H3.1 is being completely evicted and the genetic program of the cells is changing, though that aspect is not explored in this work.

A really fascinating aspect is that H3.1 gets reincorporated after the final meristem cell division for part of the differentiation process known as endocycling. Endocycling involves making more copies of the genome without going through a cell division (and seems to be a feature of many differentiating plant cells). H3.1 is incorporated during the S phase (S for synthesis, as in DNA synthesis/copying). An endocycle is basically a cycle between G2 and S phases, skipping the other stages of the cell cycle, G1 and actual mitosis or cell division. However, endocycles stop too, and indeed, at the end of endocycling ahead of a fully differentiated root cell, H3.1 is evicted again. Thus H3.1 seems to be a marker of cycling cells- either mitotic or endocycling. This dynamic seems to exist in other plant tissues as well (guard cell differentiation for instance).
More to the story
Both of these studies show two ways plant cells can determine their identity and behavior. That said, there are bigger networks of genes than the ones mentioned here. There are a lot of transcription factors that integrate cues from plant growth hormones, environmental cues, and from signals from neighboring cells (or even further away cells). Some transcription factors actually move between one cell file and another as one example of how plant cells talk to each other. Histone dynamics are also not just down to one being incorporated into the histone octamer that wraps DNA. Histones also can be marked in various ways creating different accessibility to DNA for other proteins that would bind to a region of DNA (for instance, transcription factors), this is part of epigenetic regulation.
These genetic programs are operating in all plants, creating the rich network of a plant’s inner life below and above ground.
Plant cells have different genetic programs to specify cell function. Some factors activate genes, others repress gene (or protein) function. There can be hundreds of genes involved and plant scientists are really starting to appreciate the network aspects of how genes regulate plant growth and development. These genetic programs are operating in all plants, creating the rich network of a plant’s inner life below and above ground.
References
- Rodriguez, R.E., Ercoli M.F., Debernardi J.M., Breakfield N.W., Mechhia M.A., Sabatini, M., Cools, T., De Veydler, L., Benfey, P.N., Palatnik, J.F. 2016. MicroRNA miR396 Regulates the Switch between Stem Cells and Transit-Amplifying Cells in Arabidopsis Roots. Plant Cell 27: 3354-3366 doi: www.plantcell.org/cgi/doi/10.1105/tpc.15.00452
- Otero, S., Desvoyes, B., Peiró, R., Gutierrez, C. 2016. Histone H3 Dynamics Reveal Domains with Distinct Proliferation Potential in the Arabidopsis Root. Plant Cell 28: 1361-1371 doi: www.plantcell.org/cgi/doi/10.1105/tpc.15.01003
Amazing! I read the first reference paper, but didn’t pay much attention on the second one.
LikeLike