Say you write something and submit it for editing. Your editor sends it back with comments. And you create a new version and re-submit and so on. it’s a feedback loop. With editor/writer, it’s a loop of iterative improvement resulting in something publishable (ideally).
Feedback loops in biology are also quite common.
A recent paper from Lauressergues et al. (1) in Nature highlights one such loop discovered in plants that may well exist in other organisms as well. It has to do with regulating one form of ribo-nucleic acid (RNA), one of the big 4 molecules required for life as we know it– DNA, RNA, proteins, and lipids.
RNA is funny stuff. It’s been surprising scientists for decades. It’s gone from mere conveyor of information to central player in many cellular processes.
RNA is funny stuff. It’s been surprising scientists for decades. It’s gone from mere conveyor of information to central player in many cellular processes. RNA may also be the original “molecule of life”, an early molecule to store copyable, heritable, information. It could be passed on to later generations. RNA can also carry out some chemical reactions.
After the double helix structure of DNA was discovered by Franklin et. al. in the 1950’s, the race was on to figure out just how life accessed the information stored in DNA. DNA is the molecule many familiar with as the carrier of heritable information. As it turns out, there are enzymes (one kind of protein) in cells that use one strand of DNA and turn it into the related molecule, RNA. Some RNA then gets translated (literally…there’s a code!) into proteins that then go off and do their myriad jobs, creating life, including that enzyme that makes RNA based on the DNA template. That’s mind blowing enough…but RNA does so much more.
The cellular machinery that turns RNA into protein is also made largely of RNA and it is RNA that does the protein making itself.
Some viruses have genomes that are RNA based, and some even encode enzymes able to go the other way, using an RNA template to make DNA.
In fact, biologists suspect that RNA was the first molecule of life and that DNA and proteins are essentially spin offs of what used to be an RNA world. DNA is more stable than RNA. Proteins are more flexible in what they can do; much more than just modify RNA and DNA, but also enable metabolism and complex chemsitries cells contain. So there are reasons other molecules exist, but RNA can do lots of stuff.
Hints of more
25 years ago this month, Richard Jorgensen, Antoine Stuitje and colleagues published some of their research on petunias(2,3). They were over-expressing an enzyme involved in making anthocyanin, a purple plant pigment. Their interest was in studying whether the particular enzyme, chalcone synthase, was the enzyme in making anthocyanin that was. Anthocyanins are interesting in their own right, and are what’s behind the color changing petunia I wrote about a few weeks ago. The researchers were able to make a quite purple petunia. But when Jorgensesn looked at some of his chalcone synthase over-producing petunias in the next generation, they weren’t just all purple. Some had patterns of purple and white, some were all white, and yet all still had the piece of DNA over-expressing the enzyme in their genome as well as the normal. What was going on? The scientists figured out that over-expression of the enzyme was causing silencing of the gene in some cases, as in, the RNA was no longer being produced. The over-expressing transgene has to over-produce RNA to see the silencing effect.
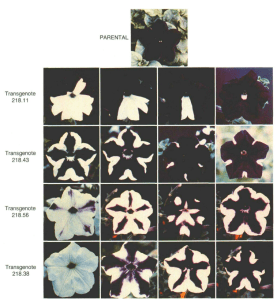
What was going on? RNA seemed to be causing a silencing effect on gene expression. Turning up the level of RNA led to a negative feedback loop.
A few years later, the team of Fire and Mello, working with worms, discovered part of what was going on. It turns out many organisms have a system designed to silence gene expression based on double stranded RNA. It turns out that long strands of double stranded RNA (dsRNA) gets chopped up into ~20 nucleotide (nt) pieces (nucleotides are the units of DNA and RNA). These 20nt RNA molecules can recognize other RNA with similar sequences and chop it up to destroy it or stop it from being made into protein. If that piece of RNA came from a DNA template, that gene could be turned off, silenced. Subsequent work in many systems and by many scientists further defined just how this system works. And discovered wrinkles in how each organism utilizes and processes its small RNAs.
Although dsRNA triggers the silencing system best, an over-abundance of single stranded RNA induce a similar effect.
One thing that was discovered is that the genome, the DNA, has sequences that get made into small RNAs. These RNAs most often act to turn off the expression of a target gene. These RNAs end up as the ~20 nucleotide molecules, but start out as bigger RNA molecules that undergo processing to its final functional form. These genes are called microRNAs, or miRNAs.
It’s one of the things behind the idea that regions of the genome not coding for proteins are not really made of “junk”.
Small proteins and small RNA expressions
Lauressergues et al. found that some pre-processed miRNA (pri-miRNAs) have regions that code for small proteins that promote the expression of that same miRNA. They dub them miPEPs, for microRNA peptide (a peptide is a string of amino acids– the constituents of proteins) in the paper. I sarcastically called them partially coding non-coding RNAs (pcncRNAs) on Twitter. Small peptides are not uncommon in biology. Insulin is an example (http://en.wikipedia.org/wiki/Insulin). It’s a small peptide that regulates blood sugar in animals and binds to a protein receptor to effect metabolism. Insulin, however, starts out as a larger protein that the insulin peptide is excised from; it does not start out small. In this case, the small peptide seems to bind to the DNA of the miRNA gene encoding it and cause more of it to be produced. It’s a positive feedback loop.
In this case, the small peptide seems to bind to the DNA of the miRNA gene encoding it and cause more of it to be produced. It’s a positive feedback loop.
The researchers showed that these small proteins are made in the plant. That miPEP expression overlaps with where the miRNA genes are active in development, and that the loss of the small peptide results in less of the final miRNA. They also show that just treating a plant with the peptide can get it to produce more of this miRNA. This may lead to some clever applications of small peptides to manipulate gene expression in plants. There are many questions that remain. As the authors say at the end of their paper:
“The discovery of miPEPs encoded by pri-miRNAs and their effect on pri-miRNA transcription raises several questions. Not least, how is pri-miRNA translation in the cytoplasm co-ordinated[sic] with the maturation of pri-miRNAs in the nucleus? How to miPEPs interact with the transcriptional machinery and what is the molecular basis of their specificity for their corresponding pri-miRNAs?”(1)
And though I am handwaving here, small RNAs exist in bacteria too as do small proteins(4) that seem to serve regulatory functions. It would not shock me to learn that small peptides coded by RNA are ancient, perhaps the first proteins existed make more RNA and these small peptides somehow helped in that function. Just how life went from an RNA world to a DNA-RNA-protein world is still a question, though there are ideas about how this complexity arose.
This paper is yet another in a long line of plant biology uncovering some new aspect of biology. There is a lot going on inside living organisms, including plants (I underscore this as it can be hard to see how a plant is like a human). This miPEP story is just the latest uncovered by scientists making careful observations and testing hypotheses.
References:
1. Lauressergues D, Couzigou JM, San Clemente H, Martinez Y, Dunand C, Becard G, Combier JP. 2015. Primary transcripts of microRNAs encode regulatory peptides. Nature. doi 10.1038/nature14346.
2. van der Krol AR, Mur AL, Beld M, Moi JNM, and Stuitje AR. 1990. Flavonoid Genes in Petunia: Addition of a Limited Number of Gene Copies May Lead to a Suppression of Gene Expression. Plant Cell 2:291-299.
3. Napoli C, Lemieux C, Jorgensen, R. 1990. lntroduction of a Chimeric Chalcone Synthase Gene into Petunia Results in Reversible Co-Suppression of Homologous Genes Ín trans. Plant Cell 2:279-289.
4. Storz, G, Wolf YI, Ramammurthi K. 2014. Small Proteins Can No Longer Be Ignored. Annu. Rev. Biochem. 83:753-77.
5 thoughts on “Small surprises and a new feedback loop.”